Neural networks, a type of computing system loosely modeled on the organization of the human brain, form the basis of many artificial intelligence systems for applications such speech recognition, computer vision, and medical image analysis.
In the field of neuroscience, researchers often use neural networks to try to model the same kind of tasks that the brain performs, in hopes that the models could suggest new hypotheses regarding how the brain itself performs those tasks. However, a group of researchers at MIT is urging that more caution should be taken when interpreting these models.
In an analysis of more than 11,000 neural networks that were trained to simulate the function of grid cells — key components of the brain’s navigation system — the researchers found that neural networks only produced grid-cell-like activity when they were given very specific constraints that are not found in biological systems.
“What this suggests is that in order to obtain a result with grid cells, the researchers training the models needed to bake in those results with specific, biologically implausible implementation choices,” says Rylan Schaeffer, a former senior research associate at MIT.
Without those constraints, the MIT team found that very few neural networks generated grid-cell-like activity, suggesting that these models do not necessarily generate useful predictions of how the brain works.
Schaeffer, who is now a graduate student in computer science at Stanford University, is the lead author of the new study, which will be presented at the 2022 Conference on Neural Information Processing Systems this month. Ila Fiete, a professor of brain and cognitive sciences and a member of MIT’s McGovern Institute for Brain Research, is the senior author of the paper. Mikail Khona, an MIT graduate student in physics, is also an author.
Modeling grid cells
Neural networks, which researchers have been using for decades to perform a variety of computational tasks, consist of thousands or millions of processing units connected to each other. Each node has connections of varying strengths to other nodes in the network. As the network analyzes huge amounts of data, the strengths of those connections change as the network learns to perform the desired task.
In this study, the researchers focused on neural networks that have been developed to mimic the function of the brain’s grid cells, which are found in the entorhinal cortex of the mammalian brain. Together with place cells, found in the hippocampus, grid cells form a brain circuit that helps animals know where they are and how to navigate to a different location.
Place cells have been shown to fire whenever an animal is in a specific location, and each place cell may respond to more than one location. Grid cells, on the other hand, work very differently. As an animal moves through a space such as a room, grid cells fire only when the animal is at one of the vertices of a triangular lattice. Different groups of grid cells create lattices of slightly different dimensions, which overlap each other. This allows grid cells to encode a large number of unique positions using a relatively small number of cells.
This type of location encoding also makes it possible to predict an animal’s next location based on a given starting point and a velocity. In several recent studies, researchers have trained neural networks to perform this same task, which is known as path integration.
To train neural networks to perform this task, researchers feed into it a starting point and a velocity that varies over time. The model essentially mimics the activity of an animal roaming through a space, and calculates updated positions as it moves. As the model performs the task, the activity patterns of different units within the network can be measured. Each unit’s activity can be represented as a firing pattern, similar to the firing patterns of neurons in the brain.
In several previous studies, researchers have reported that their models produced units with activity patterns that closely mimic the firing patterns of grid cells. These studies concluded that grid-cell-like representations would naturally emerge in any neural network trained to perform the path integration task.
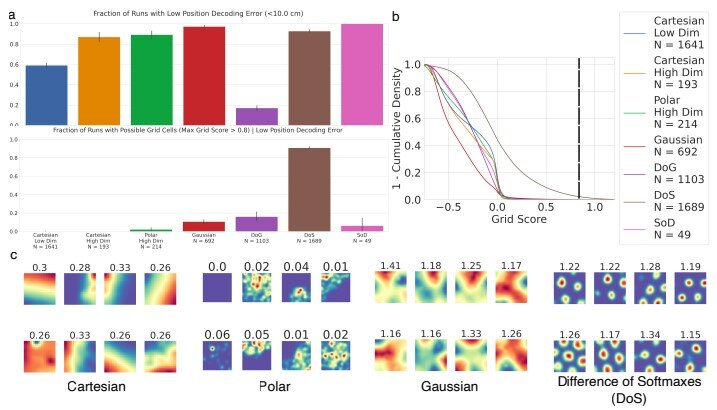
However, the MIT researchers found very different results. In an analysis of more than 11,000 neural networks that they trained on path integration, they found that while nearly 90 percent of them learned the task successfully, only about 10 percent of those networks generated activity patterns that could be classified as grid-cell-like. That includes networks in which even only a single unit achieved a high grid score.
The earlier studies were more likely to generate grid-cell-like activity only because of the constraints that researchers build into those models, according to the MIT team.
“Earlier studies have presented this story that if you train networks to path integrate, you're going to get grid cells. What we found is that instead, you have to make this long sequence of choices of parameters, which we know are inconsistent with the biology, and then in a small sliver of those parameters, you will get the desired result,” Schaeffer says.
More biological models
One of the constraints found in earlier studies is that the researchers required the model to convert velocity into a unique position, reported by one network unit that corresponds to a place cell. For this to happen, the researchers also required that each place cell correspond to only one location, which is not how biological place cells work: Studies have shown that place cells in the hippocampus can respond to up to 20 different locations, not just one.
When the MIT team adjusted the models so that place cells were more like biological place cells, the models were still able to perform the path integration task, but they no longer produced grid-cell-like activity. Grid-cell-like activity also disappeared when the researchers instructed the models to generate different types of location output, such as location on a grid with X and Y axes, or location as a distance and angle relative to a home point.
“If the only thing that you ask this network to do is path integrate, and you impose a set of very specific, not physiological requirements on the readout unit, then it's possible to obtain grid cells,” Fiete says. “But if you relax any of these aspects of this readout unit, that strongly degrades the ability of the network to produce grid cells. In fact, usually they don't, even though they still solve the path integration task.”
Therefore, if the researchers hadn’t already known of the existence of grid cells, and guided the model to produce them, it would be very unlikely for them to appear as a natural consequence of the model training.
The researchers say that their findings suggest that more caution is warranted when interpreting neural network models of the brain.
“When you use deep learning models, they can be a powerful tool, but one has to be very circumspect in interpreting them and in determining whether they are truly making de novo predictions, or even shedding light on what it is that the brain is optimizing,” Fiete says.
Kenneth Harris, a professor of quantitative neuroscience at University College London, says he hopes the new study will encourage neuroscientists to be more careful when stating what can be shown by analogies between neural networks and the brain.
“Neural networks can be a useful source of predictions. If you want to learn how the brain solves a computation, you can train a network to perform it, then test the hypothesis that the brain works the same way. Whether the hypothesis is confirmed or not, you will learn something,” says Harris, who was not involved in the study. “This paper shows that ‘postdiction’ is less powerful: Neural networks have many parameters, so getting them to replicate an existing result is not as surprising.”
When using these models to make predictions about how the brain works, it’s important to take into account realistic, known biological constraints when building the models, the MIT researchers say. They are now working on models of grid cells that they hope will generate more accurate predictions of how grid cells in the brain work.
“Deep learning models will give us insight about the brain, but only after you inject a lot of biological knowledge into the model,” Khona says. “If you use the correct constraints, then the models can give you a brain-like solution.”
The research was funded by the Office of Naval Research, the National Science Foundation, the Simons Foundation through the Simons Collaboration on the Global Brain, and the Howard Hughes Medical Institute through the Faculty Scholars Program. Mikail Khona was supported by the MathWorks Science Fellowship.